
In human genetics, a human mitochondrial DNA haplogroup is a haplogroup defined by differences in human mitochondrial DNA. Haplogroups are used to represent the major branch points on the mitochondrial phylogenetic tree. Understanding the evolutionary path of the female lineage has helped population geneticists trace the matrilineal inheritance of modern humans back to human origins in Africa and the subsequent spread around the globe.
The letter names of the haplogroups (not just mitochondrial DNA haplogroups) run from A to Z. As haplogroups were named in the order of their discovery, the alphabetical ordering does not have any meaning in terms of actual genetic relationships.
The hypothetical woman at the root of all these groups (meaning just the mitochondrial DNA haplogroups) is the matrilineal most recent common ancestor (MRCA) for all currently living humans. She is commonly called Mitochondrial Eve.
The rate at which mitochondrial DNA mutates is known as the mitochondrial molecular clock. It is an area of ongoing research with one study reporting one mutation per 8000 years.[2]
YouTube Encyclopedic
-
1/5Views:496 73010 838110 817295 032105 043
-
mtDNA shows how humans migrated across the World
-
Where did the mtDNA haplogroups live?
-
Migration path of Y-chromosome Adam
-
Haplogroup Map of the World: Your Genetic Surname (+Download Link)
-
Mother of Humanity (Mitochondrial Eve)
Transcription
Phylogeny
| Part of a series on |
| Genetic genealogy |
|---|
| Concepts |
| Related topics |

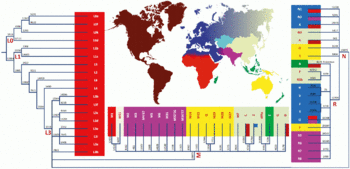
This phylogenetic tree is based Van Oven (2009).[4] In June 2022, an alternative phylogeny for haplogroup L was suggested[5]
Major mtDNA Haplogroups

Macro-haplogroup L
Macro-haplogroup L is the most basal of human mtDNA haplogroups, from which all other haplogroups descend (specifically, from haplogroup L3). It is found mostly in Africa.
- Haplogroup L0
- L1-7
- Haplogroup L1
- L2-7
- L3'4'6
- L5'7
Macro-haplogroup M
Macro-haplogroup M is found mostly in Asia and the Americas. Its descendants are haplogroup M, haplogroup C, haplogroup Z, haplogroup D, haplogroup E, haplogroup G and haplogroup Q.
Macro-haplogroup N
Macro-haplogroup N is found mostly in Australia, the Americas and parts of Asia. Its descendants are haplogroup N, haplogroup O, haplogroup A, haplogroup S, haplogroup I, haplogroup W, haplogroup X and haplogroup Y, as well as macro-haplogroup R.
Macro-haplogroup R
Macro-haplogroup R is found mostly in Europe, Northern Africa, the Pacific and parts of Asia and the Americas. Its descendants are haplogroup R, haplogroup B, haplogroup F, haplogroup H, haplogroup V, haplogroup J, haplogroup T, haplogroup U and haplogroup K
Chronology
| Haplogroup | Est. time of origin (kya)[6] | Possible place of origin | Highest frequencies |
|---|---|---|---|
| L | 200 | Africa | |
| L1-6 | 170 | East Africa | |
| L2-6 | 150 | East Africa | |
| L0 | 150 | East Africa | |
| L1 | 140 | Central Africa | |
| L3-6 | 130 | ||
| L5 | 120 | ||
| L2 | 90 | ||
| L3 | 70 | East Africa | |
| N | 70 | East Africa or West Asia | |
| M | 60 | East Africa, West Asia or South Asia | |
| R | 60 | South Asia or Southeast Asia | |
| U | 55 | North-East Africa or India (South Asia) | |
| RT'JT | 55 | Middle East | |
| JT | 50 | Middle East | |
| U8 | 50 | Western Asia | |
| R9 | 47 | ||
| B4 | 44 | ||
| F | 43 | ||
| U4'9 | 42 | Central Asia | |
| U5 | 35 | Western Asia | |
| U6 | 35 | North Africa | |
| J | 35 | ||
| X | 30 | ||
| K | 30 | ||
| U5a | 27 | ||
| HV | 27 | Near East | |
| J1a | 27 | Near East | |
| T | 27 | Mesopotamia | |
| K1 | 27 | ||
| I | 26 | ||
| J1 | 24 | Near East | |
| W | 20 | ||
| U4 | 20 | Central Asia | |
| X2 | 20 | ||
| H | 20 | Western Asia | |
| U5a1 | 18 | Europe | |
| J1b | 11 | ||
| V | 14 | ||
| X2a | 13 | North America | |
| H1 | 12 | ||
| H3 | 12 | ||
| X1 | 10 |
Geographical distribution
A 2004 paper suggested that the haplogroups most common in modern West Asian, North African and European populations were: H, J, K, N1, T, U4, U5, V, X and W.[7]
African haplogroups: L0, L1, L2, L3, L4, L5, L6, T, U5a
Australian haplogroups: M42a, M42c, M14, M15, Q, S, O, N, P. (Refs 1, 2, 3, 4, 5, 6)
Asian haplogroups: F, C, W, M, D, N, K, U, T, A, B, C, Z, U many number variants to each section
See also
- Human mitochondrial genetics
- Genetic genealogy
- Matrilineality
- Human Y-chromosome DNA haplogroups
- Population genetics
References
- ^ Rishishwar L, Jordan IK (2017). "Implications of human evolution and admixture for mitochondrial replacement therapy". BMC Genomics. 18 (1): 140. doi:10.1186/s12864-017-3539-3. PMC 5299762. PMID 28178941.
- ^ Loogvali, Eva-Liis; Kivisild, Toomas; Margus, Tõnu; Villems, Richard (2009), O'Rourke, Dennis (ed.), "Explaining the Imperfection of the Molecular Clock of Hominid Mitochondria", PLOS ONE, 4 (12): e8260, Bibcode:2009PLoSO...4.8260L, doi:10.1371/journal.pone.0008260, PMC 2794369, PMID 20041137
- ^ Kivisild T (2015). "Maternal ancestry and population history from whole mitochondrial genomes". Investig Genet. 6: 3. doi:10.1186/s13323-015-0022-2. PMC 4367903. PMID 25798216.
- ^ a b van Oven M, Kayser M (February 2009). "Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation". Human Mutation. 30 (2): E386–94. doi:10.1002/humu.20921. PMID 18853457. S2CID 27566749.
- ^ Maier P, Runfeldt G, Estes R, Vilar M (2022). "African mitochondrial haplogroup L7: a 100,000-year-old maternal human lineage discovered through reassessment and new sequencing". Nature. 12 (1): 10747. Bibcode:2022NatSR..1210747M. doi:10.1038/s41598-022-13856-0. PMC 9232647. PMID 35750688. S2CID 250021505.
- ^ "Correcting for Purifying Selection: An Improved Human Mitochondrial Molecular Clock Supplementary" (PDF). 2009: 82–83 [89]. Archived from the original (PDF) on 2009-12-29.
{{cite journal}}: Cite journal requires|journal=(help) - ^ Villems, Richard; Usanga, Esien; Mikerezi, Ilia; Gölge, Mukaddes; Claustres, Mireille; Michalodimitrakis, Emmanuel N.; Pappa, Kalliopi I.; Anagnou, Nicholas P.; Chaventré, André; Moisan, Jean-Paul; Richard, Christelle; Grechanina, Elena; Balanovska, Elena V.; Rudan, Pavao; Puzyrev, Valery; Stepanov, Vadim; Khusnutdinova, Elsa K.; Gusar, Vladislava; Balanovsky, Oleg P.; Peričić, Marijana; Barać, Lovorka; Golubenko, Maria; Lunkina, Arina; Laos, Sirle; Pennarun, Erwan; Parik, Jüri; Tolk, Helle-Viivi; Reidla, Maere; Tambets, Kristiina; Metspalu, Ene; Kivisild, Toomas; Derenko, Miroslava V.; Malyarchuk, Boris A.; Roostalu, Urmas; Loogväli, Eva-Liis (November 1, 2004). "Disuniting Uniformity: A Pied Cladistic Canvas of mtDNA Haplogroup H in Eurasia". Molecular Biology and Evolution. 21 (11): 2012–2021. doi:10.1093/molbev/msh209. PMID 15254257 – via academic.oup.com.
External links
- Mitochondrial phylogenetic trees
- Mannis van Oven's PhyloTree.org
- PhyloD3 – D3.js-based phylogenetic tree based on PhyloTree
- Mitochondrial haplogroup skeleton
- Vincent Macaulay's Mitochondrial haplogroup motifs
- List of mtDNA haplogroup projects
- MitoTool: a web server for the analysis and retrieval of human mitochondrial DNA sequence variations
- HaploGrep: mtDNA haplogroup determination based on PhyloTree.org
- HaploFind – fast automatic haplogroup assignment pipeline for human mitochondrial DNA Archived 2016-06-11 at the Wayback Machine
|
Phylogenetic tree of human mitochondrial DNA (mtDNA) haplogroups | |||||||||||||||||||||||||||||||||||||||
| Mitochondrial Eve (L) | |||||||||||||||||||||||||||||||||||||||
| L0 | L1–6 | ||||||||||||||||||||||||||||||||||||||
| L1 | L2 | L3 | L4 | L5 | L6 | ||||||||||||||||||||||||||||||||||
| M | N | ||||||||||||||||||||||||||||||||||||||
| CZ | D | E | G | Q | O | A | S | R | I | W | X | Y | |||||||||||||||||||||||||||
| C | Z | B | F | R0 | pre-JT | P | U | ||||||||||||||||||||||||||||||||
| HV | JT | K | |||||||||||||||||||||||||||||||||||||
| H | V | J | T | ||||||||||||||||||||||||||||||||||||
