
Protein electrophoresis is a method for analysing the proteins in a fluid or an extract. The electrophoresis may be performed with a small volume of sample in a number of alternative ways with or without a supporting medium, namely agarose or polyacrylamide. Variants of gel electrophoresis include SDS-PAGE, free-flow electrophoresis, electrofocusing, isotachophoresis, affinity electrophoresis, immunoelectrophoresis, counterelectrophoresis, and capillary electrophoresis. Each variant has many subtypes with individual advantages and limitations. Gel electrophoresis is often performed in combination with electroblotting or immunoblotting to give additional information about a specific protein.[1]
YouTube Encyclopedic
-
1/5Views:835 023224 6441 102 733192 69880 664
-
Gel electrophoresis | Chemical processes | MCAT | Khan Academy
-
SDS-PAGE, Sodium Dodecyl Sulfate–PolyAcrylamide Gel Electrophoresis–Animation
-
Gel Electrophoresis
-
Polyacrylamide Gel Electrophoresis- PAGE - Amrita University
-
SDS PAGE | polyacrylamide gel electrophoresis
Transcription
Today, we'll be talking about gel electrophoresis. What is gel electrophoresis, you might ask. Well, it's a lab technique usually used in the biochemistry lab for separating out DNA or proteins based on their size. And let's talk about how it works. So first, you need to have the gel. This can be made out of different kinds of substances, such as agarose and polyacrylamide, both of which I'll discuss later. And the electrophoresis part of it means that you need to have an electrical field passing through the gel to get the bands to move. So to create an electrical field, we have to have a cathode and an anode. The cathode is on this side. Remember from general chemistry that at the cathode, reduction takes place, so you'll have a negative charge. But on the other side, where the anode is, you'll have a positive charge, because this is where oxidation is taking place. And to power this, you'll need some kind of battery to connect them. And these are both usually also connected to a big box that contains the gel electrophoresis apparatus. In order for charge to flow across, you need something that will conduct electricity, so you have a buffer that's usually composed of different ions. So this is completely covering this entire gel. And remember that it's always there, but I'm going to erase it just because it's going to get a little messy for what I need to show you next. So next, you'll load a sample of DNA. In order to load your sample, you'll need to take a pipette and put it into one if these wells. So usually you'll want to make sure that you have a dye mixed in with your sample so that you can see it as it's running. So I'll show that as a pink line here, so I've filled that well, followed by yellow here, and green here. So that's just the color of the dye. It can really be any color you want. This will just help you track the movement of the bands later on. Once you have these bands, what will happen once you turn on the electrical field? Remember that DNA has a negatively charged backbone because of all the phosphate groups, so what you'll actually observe is that they'll travel towards the positive end. So maybe after a short period of time, what you'll see is that-- it'll look something like this. Maybe the pink will have split up into a few bands, indicating that there's a few different sized fragments in there. With the yellow, you might only see one band. And with the green, you might actually have two bands, but they're so close together still that it's kind of hard to tell. So how can you really see what's going on? The solution is to let this run for a longer period of time. And eventually, what you'll see is shown here at the bottom. You'll see that whatever sized fragments were in your original samples have effectively split up by their size. And note that in pink, you'll always want to load something known as a DNA ladder. A DNA ladder is like a standard. This is something that you buy from a company, and they tell you exactly what sizes their fragments so that you can match them up to your unknown samples. So this could be 400 base pairs, 200 base pairs, and 100 base pairs. And as you'll note, the smallest fragments travel the furthest. This is because the smallest things are really easy to push with the electrical field. But when you have such a big molecule, or rather a big DNA fragment, it can be hard to move. So you can see that the 400 base pair doesn't move too fast or too far. And what's this tell us about our unknown yellow and green samples? This shows us that the yellow sample has a band that's 200 base pairs long. And the green sample actually was composed of two different fragments, one that was 100 base pairs and another that was 200 base pairs. So what would we do with this information? If you needed a particular size of DNA, say for the next step of your experiment, if you wanted to insert it into a plasmid or a vector, you could cut this out of the gel and use it for that. Now let's talk about the two kinds of gels that are most commonly used. The first is agarose, and the second is SDS-PAGE. So agarose is a gel that's usually used for separating big pieces of DNA. So if you think about the pore size in the agarose, it has pretty big pores, so imagine it looking kind of like this. The gel is pretty big. There's big holes here, so that you'll be able to separate out the big pieces of DNA that come through. However, if you're trying to separate out little pieces, it won't be that obvious, because they'll all just race through these giant holes. So remember that this is for big DNA fragments. Usually, this is for DNA that's bigger than 50 base pairs. SDS-PAGE, on the other hand, can be used for very small things. So imagine that being a much finer weaving with smaller pores. Although this can be used for small pieces of DNA, it can also be used for proteins. You might be wondering, what does SDS-PAGE even stand for. The SDS part is Sodium Dodecyl Sulfate. This is a chemical agent that denatures proteins, disrupting any non-covalent interactions they may have. This makes it so that the charge of the proteins isn't a factor when they're separating out onto the gel, and they're only being separated strictly by size. The PAGE part is PolyAcrylamide Gel Electrophoresis, or we'll just leave it at GE. So polyacrylamide is the substance that gel's made out. So how can we remember the difference between these two types of gels? Remember that SDS-PAGE is for small DNA or protein cells. S for small, and S for SDS. And agarose is for bigger fragments of DNA. So today we've talked about how you would setup a gel electrophoresis, why it works, and how you would want to pick the substance but your gel's made of. If you were really doing this in the lab, now that you have your fragments of known size of DNA or protein, you could either sequence them or use them in other molecular techniques.
Denaturing gel methods
SDS-PAGE
SDS-PAGE, sodium dodecyl sulfate polyacrylamide gel electrophoresis, describes a collection of related techniques to separate proteins according to their electrophoretic mobility (a function of the molecular weight of a polypeptide chain) while in the denatured (unfolded) state. In most proteins, the binding of SDS to the polypeptide chain imparts an even distribution of charge per unit mass, thereby resulting in a fractionation by approximate size during electrophoresis.[2]
SDS is a strong detergent agent used to denature native proteins to unfolded, individual polypeptides. When a protein mixture is heated to 100 °C in presence of SDS, the detergent wraps around the polypeptide backbone. In this process, the intrinsic charges of polypeptides becomes negligible when compared to the negative charges contributed by SDS. Thus polypeptides after treatment become rod-like structures possessing a uniform charge density, that is same net negative charge per unit length. The electrophoretic mobilities of these proteins will be a linear function of the logarithms of their molecular weights.[3]
Native gel methods
Native gels, also known as non-denaturing gels, analyze proteins that are still in their folded state. Thus, the electrophoretic mobility depends not only on the charge-to-mass ratio, but also on the physical shape and size of the protein.[4]
Blue native PAGE
BN-PAGE is a native PAGE technique, where the Coomassie brilliant blue dye provides the necessary charges to the protein complexes for the electrophoretic separation.[5][6] The disadvantage of Coomassie is that in binding to proteins it can act like a detergent causing complexes to dissociate. Another drawback is the potential quenching of chemoluminescence (e.g. in subsequent western blot detection or activity assays) or fluorescence of proteins with prosthetic groups (e.g. heme or chlorophyll) or labelled with fluorescent dyes.[citation needed]
Clear native PAGE
CN-PAGE (commonly referred to as Native PAGE) separates acidic water-soluble and membrane proteins in a polyacrylamide gradient gel. It uses no charged dye so the electrophoretic mobility of proteins in CN-PAGE (in contrast to the charge shift technique BN-PAGE) is related to the intrinsic charge of the proteins.[7] The migration distance depends on the protein charge, its size and the pore size of the gel. In many cases this method has lower resolution than BN-PAGE, but CN-PAGE offers advantages whenever Coomassie dye would interfere with further analytical techniques, for example it has been described as a very efficient microscale separation technique for FRET analyses.[8] Additionally, as CN-PAGE does not require the harsh conditions of BN-PAGE, it can retain the supramolecular assemblies of membrane protein complexes that would be dissociated in BN-PAGE.[7]
Preparative native PAGE
The folded protein complexes of interest separate cleanly and predictably without the risk of denaturation due to the specific properties of the polyacrylamide gel, electrophoresis buffer solution, electrophoretic equipment and standardized parameters used. The separated proteins are continuously eluted into a physiological eluent and transported to a fraction collector. In four to five PAGE fractions each the different metal cofactors can be identified and absolutely quantified by high-resolution ICP-MS. The associated structures of the isolated metalloproteins in these fractions can be specifically determined by solution NMR spectroscopy.[9]
Buffer systems

Most protein separations are performed using a "discontinuous" (or DISC) buffer system that significantly enhances the sharpness of the bands within the gel. During electrophoresis in a discontinuous gel system, an ion gradient is formed in the early stage of electrophoresis that causes all of the proteins to focus into a single sharp band. The formation of the ion gradient is achieved by choosing a pH value at which the ions of the buffer are only moderately charged compared to the SDS-coated proteins. These conditions provide an environment in which Kohlrausch's reactions determine the molar conductivity. As a result, SDS-coated proteins are concentrated to several fold in a thin zone of the order of 19 μm within a few minutes. At this stage all proteins migrate at the same migration speed by isotachophoresis. This occurs in a region of the gel that has larger pores so that the gel matrix does not retard the migration during the focusing or "stacking" event.[10][11] Separation of the proteins by size is achieved in the lower, "resolving" region of the gel. The resolving gel typically has a much smaller pore size, which leads to a sieving effect that now determines the electrophoretic mobility of the proteins. At the same time, the separating part of the gel also has a pH value in which the buffer ions on average carry a greater charge, causing them to "outrun" the SDS-covered proteins and eliminate the ion gradient and thereby the stacking effect.[citation needed]
A very widespread discontinuous buffer system is the tris-glycine or "Laemmli" system that stacks at a pH of 6.8 and resolves at a pH of ~8.3-9.0. A drawback of this system is that these pH values may promote disulfide bond formation between cysteine residues in the proteins because the pKa of cysteine ranges from 8-9 and because reducing agent present in the loading buffer doesn't co-migrate with the proteins. Recent advances in buffering technology alleviate this problem by resolving the proteins at a pH well below the pKa of cysteine (e.g., bis-tris, pH 6.5) and include reducing agents (e.g. sodium bisulfite) that move into the gel ahead of the proteins to maintain a reducing environment. An additional benefit of using buffers with lower pH values is that the acrylamide gel is more stable at lower pH values, so the gels can be stored for long periods of time before use.[12][13]
SDS gradient gel electrophoresis of proteins
As voltage is applied, the anions (and negatively charged sample molecules) migrate toward the positive electrode (anode) in the lower chamber, the leading ion is Cl− ( high mobility and high concentration); glycinate is the trailing ion (low mobility and low concentration). SDS-protein particles do not migrate freely at the border between the Cl− of the gel buffer and the Gly− of the cathode buffer. Friedrich Kohlrausch found that Ohm's law also applies to dissolved electrolytes. Because of the voltage drop between the Cl− and Glycine-buffers, proteins are compressed (stacked) into micrometer thin layers.[14] The boundary moves through a pore gradient and the protein stack gradually disperses due to a frictional resistance increase of the gel matrix. Stacking and unstacking occurs continuously in the gradient gel, for every protein at a different position. For a complete protein unstacking the polyacrylamide-gel concentration must exceed 16% T. The two-gel system of "Laemmli" is a simple gradient gel. The pH discontinuity of the buffers is of no significance for the separation quality, and a "stacking-gel" with a different pH is not needed.[15]
Visualization
The most popular protein stain is Coomassie brilliant blue. It is an anionic dye, which non-specifically binds to proteins. Proteins in the gel are fixed by acetic acid and simultaneously stained. The excess dye incorporated into the gel can be removed by destaining with the same solution without the dye. The proteins are detected as blue bands on a clear background.[16][17]
When more sensitive method than staining by Coomassie is needed, silver staining is usually used. Silver staining is a sensitive procedure to detect trace amounts of proteins in gels, but can also visualize nucleic acid or polysaccharides.[17]
Visualization methods without using a dye such as Coomassie and silver are available on the market.[18] For example Bio-Rad Laboratories markets ”stain-free” gels for SDS-PAGE gel electrophoresis. Alternatively, reversible fluorescent dyes, such as those from Azure Biosystems such as AzureRed or Azure TotalStain Q can be used.[17][18][19]
Similarly as in nucleic acid gel electrophoresis, tracking dye is often used. Anionic dyes of a known electrophoretic mobility are usually included in the sample buffer. A very common tracking dye is Bromophenol blue. This dye is coloured at alkali and neutral pH and is a small negatively charged molecule that moves towards the anode. Being a highly mobile molecule it moves ahead of most proteins.[20]
Medical applications

In medicine, protein electrophoresis is a method of analysing the proteins mainly in blood serum. Before the widespread use of gel electrophoresis, protein electrophoresis was performed as free-flow electrophoresis (on paper) or as immunoelectrophoresis.[citation needed]
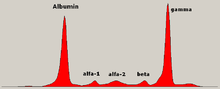
Traditionally, two classes of blood proteins are considered: serum albumin and globulin. They are generally equal in proportion, but albumin as a molecule is much smaller and lightly, negatively-charged, leading to an accumulation of albumin on the electrophoretic gel. A small band before albumin represents transthyretin (also named prealbumin). Some forms of medication or body chemicals can cause their own band, but it usually is small. Abnormal bands (spikes) are seen in monoclonal gammopathy of undetermined significance and multiple myeloma, and are useful in the diagnosis of these conditions.[citation needed]
The globulins are classified by their banding pattern (with their main representatives):[citation needed]
- The alpha (α) band consists of two parts, 1 and 2:
- α1 - α1-antitrypsin, α1-acid glycoprotein.
- α2 - haptoglobin, α2-macroglobulin, α2-antiplasmin, ceruloplasmin.
- The beta (β) band - transferrin, LDL, complement
- The gamma (γ) band - immunoglobulin (IgA, IgD, IgE, IgG and IgM). Paraproteins (in multiple myeloma) usually appear in this band.
See also
- Affinity electrophoresis
- Electroblotting
- Electrofocusing
- Fast parallel proteolysis (FASTpp)
- Gel electrophoresis
- Immunoelectrophoresis
- Immunofixation
- Native gel electrophoresis
- Paraprotein
- QPNC-PAGE
- SDD-AGE
References
- ^ Michov, Budin (2022). Electrophoresis Fundamentals: Essential Theory and Practice. De Gruyter. p. 490. doi:10.1515/9783110761641. ISBN 9783110761641. S2CID 247987700.
- ^ Stringer, R. (2005). "Electrophoresis". Encyclopedia of Analytical Science (2nd ed.). Royal Liverpool University Hospital. p. 360. doi:10.1016/B0-12-369397-7/00120-5. ISBN 978-0-12-369397-6.
- ^ Meredith, S C (1984). "The determination of molecular weight of proteins by gel permeation chromatography in organic solvents". Journal of Biological Chemistry. 259 (19): 11682–11685. doi:10.1016/s0021-9258(20)71263-9. ISSN 0021-9258. PMID 6480578.
- ^ Eubel, Holger; Braun, Hans-Peter; Millar, AHarvey (2005). "Blue-native PAGE in plants: a tool in analysis of protein-protein interactions". Plant Methods. 1 (1): 11. doi:10.1186/1746-4811-1-11. ISSN 1746-4811. PMC 1308860. PMID 16287510.
- ^ Schägger, Hermann; von Jagow, Gebhard (1991). "Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form". Analytical Biochemistry. 199 (2): 223–231. doi:10.1016/0003-2697(91)90094-A. PMID 1812789.
- ^ Wittig, Ilka; Braun, Hans-Peter; Schägger, Hermann (2006). "Blue native PAGE". Nat. Protoc. 1 (1): 418–428. doi:10.1038/nprot.2006.62. PMID 17406264. S2CID 19715017.
- ^ a b Wittig, IIlka; Schägger, Hermann (2005-10-11). "Advantages and limitations of clear-native PAGE". Proteomics. 5 (17): 4338–4346. doi:10.1002/pmic.200500081. PMID 16220535. S2CID 23396231.
- ^ Gavin, Paul D.; Devenish, Rodney J.; Prescott, Mark (2003). "FRET reveals changes in the F1–stator stalk interaction during activity of F1F0-ATP synthase". Biochim Biophys Acta. 1607 (2–3): 167–79. doi:10.1016/j.bbabio.2003.09.013. PMID 14670607.
- ^ Kastenholz, Bernd (2004). "Preparative Native Continuous Polyacrylamide Gel Electrophoresis (PNC-PAGE): An Efficient Method for Isolating Cadmium Cofactors in Biological Systems". Analytical Letters. 37 (4): 657–665. doi:10.1081/AL-120029742. ISSN 0003-2719. S2CID 97636537.
- ^ Ornstein, Leonard (December 1964). "Disc Electrophoresis. I. Background and Theory". Annals of the New York Academy of Sciences. 121 (2): 321–349. Bibcode:1964NYASA.121..321O. CiteSeerX 10.1.1.140.7598. doi:10.1111/j.1749-6632.1964.tb14207.x. PMID 14240533. S2CID 28591995.
- ^ Davis, Baruch J. (December 1964). "Disc Electrophoresis. 2, Method and application to human serum proteins". Annals of the New York Academy of Sciences. 121 (2): 404–427. Bibcode:1964NYASA.121..404D. doi:10.1111/j.1749-6632.1964.tb14213.x. PMID 14240539. S2CID 30512118.
- ^ Schägger, Hermann; von Jagow, Gebhard (1987-11-01). "Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa". Analytical Biochemistry. 166 (2): 368–379. doi:10.1016/0003-2697(87)90587-2. ISSN 0003-2697. PMID 2449095.
- ^ Wiltfang, Jens; Arold, Norbert; Neuhoff, Volker (1991). "A new multiphasic buffer system for sodium dodecyl sulfate-polyacrylamide gel electrophoresis of proteins and peptides with molecular masses 100 000–1000, and their detection with picomolar sensitivity". Electrophoresis. 12 (5): 352–366. doi:10.1002/elps.1150120507. ISSN 0173-0835. PMID 1718736. S2CID 40101706.
- ^ Kohlrausch, Friedr (1897). "Ueber Concentrations-Verschiebungen durch Electrolyse im Inneren von Lösungen und Lösungsgemischen". Annalen der Physik und Chemie. 62 (10): 209–239. Bibcode:1897AnP...298..209K. doi:10.1002/andp.18972981002.
- ^ Westermeier, Reiner (2016-05-02). Electrophoresis in Practice: Guide to Methods and Applications of DNA and Protein Separations, A (5th ed.). Wiley. p. 43. doi:10.1002/9783527695188. ISBN 978-3-527-69518-8.
- ^ Westermeier, Reiner (2016-02-26). Electrophoresis in Practice: A Guide to Methods and Applications of DNA and Protein Separations (5th ed.). Wiley. doi:10.1002/9783527695188. ISBN 978-3-527-69518-8.
- ^ a b c Sasse, Joachim; Gallagher, Sean R. (2009). "Staining Proteins in Gels". Current Protocols in Molecular Biology. 85 (1): Unit 10.6. doi:10.1002/0471142727.mb1006s85. ISSN 1934-3639. PMID 19170026. S2CID 205153300.
- ^ a b Singer, Victoria L.; Haugland, Richard P. (1999). "Fluorescent Imaging of Nucleic Acids and Proteins in Gels". Fluorescent and Luminescent Probes for Biological Activity (2nd ed.). Academic Press. pp. 58–61. doi:10.1016/b978-012447836-7/50006-3. ISBN 978-0-12-447836-7.
- ^ "In-gel Fluorescence". Azure Biosystems. Retrieved 2023-12-02.
- ^ Westermeier, Reiner (2016-05-02). Electrophoresis in Practice: Guide to Methods and Applications of DNA and Protein Separations, A (5th ed.). Wiley. pp. 2, 12, 121. doi:10.1002/9783527695188. ISBN 978-3-527-69518-8.
External links
- Educational resource for protein electrophoresis
- Gel electrophoresis of proteins Archived 2021-01-26 at the Wayback Machine
